How We Deliver: Applied Sciences for the Best of Reasons
Noblis harnesses the power of science, technology and engineering for national security and the public good.
Why Noblis for Applied Science Laboratories and Capabilities?
Our research center and applied sciences laboratories harness science-driven expertise. From genomics to nuclear physics to advance CBRNE and public health missions, Noblis answers the call for a higher standard of research and innovation.
Extended Reality (XR) Training for Increased Efficiencies and Lower Costs
See how Noblis utilizes XR, enabling trainees to build proficiency in CBRNE procedures faster while lowering labor and travel costs and reducing the logistical burdens of in-person training.

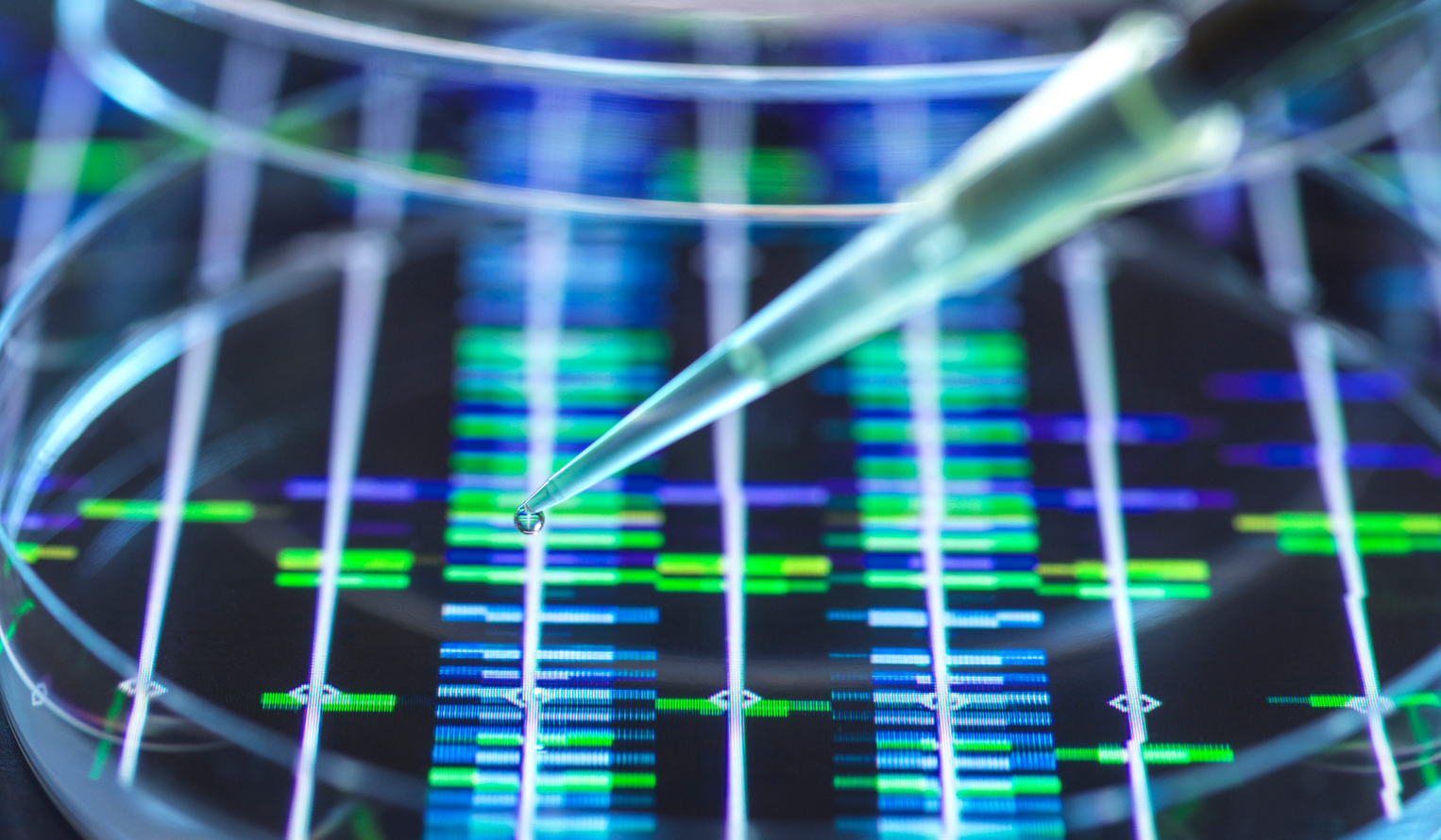
Applied DNA Sciences: Sequencing in the Field
Discover how Noblis is liberating biological threat identification from the laboratory through our applied DNA sciences capabilities. We developed a portable, field-ready DNA sequencing and analysis system that is expanding the potential of DNA analysis to improve health, science and national security.

Article: Biosurveillance has Become an Information Technology
The COVID pandemic taught us that biosurveillance and biohazard detection have entered a new phase, transformed by big data and data analytics. In effect, biosurveillance has become an information technology (IT). Noblis’ Dr. Katharine Jennings talks about molecular-based biosurveillance and how she’s been involved in building the bridge from surveillance science to IT.
Doing Business for the Best of Reasons
Hand in hand with innovation, we are committed to the highest ethical standards in all our business interactions. Discover our commitment and learn more about our code of conduct.